|
After aligning sample traces to the reference, SGCaller calculates the total area under all four peaks for the base to be genotyped. SGCaller determines each file's read direction from its name. SGCaller reads a collection of trace files in ABI or Staden (SCF) format, finds the variant position and tries to call the genotype for each, and presents them for the user to confirm and enhance the calling. We therefore developed a program to perform largely automatic assignment of SNP genotypes based on annotated SNPs in a GenBank reference sequence (SNPs annotated with the “variation” feature key file format specification provided at Sequences in the required GenBank format are available from various sources they can be generated for a locus of interest through interfaces such as ENSEMBL ( or by the user using local sequence and SNP information with tools such as the free ARTEMIS ( sequence editor. The same applies to sequencing genotypes-we think that an efficient and user-friendly way to review, assign, or correct genotype calls is essential. Finally, a moderately complex set of programs and scripts in a UNIX environment needs to be maintained for the relatively straightforward task of genotype assessment at predefined sites.ĭespite the existence of several automated methods for sequence trace analysis, sequence finishing still requires some user interaction. GenBank® files cannot be directly processed, and the incorporation of manual call correction performed in Consed requires extra effort. Because the interface is mainly designed to support contig assembly, the “vertical” assessment of hundreds of sequences at one specific base is difficult. However, PolyPhred and Consed had a different and broader design scope than that of genotype calling. PolyPhred ( 4) can automatically call genotypes at predefined sites through the use of manual tags in the “.polyphredrc” file. There is scarce supportive software (except in specialized cases) because this is an unusual and underappreciated use of sequencing technology. When “genotyping by sequencing” is performed, extracting genotype information is very cumbersome. For regions of very high variability, such as the HLA region, sequencing is an established genotyping method ( 2, 3), and specialized commercial analysis software is available. 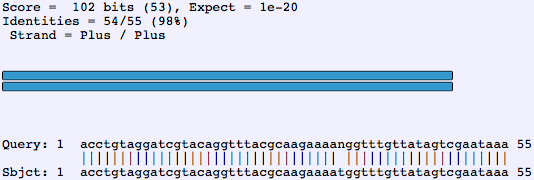
For these SNPs, researchers may be prepared to use sequencing as an alternative method for genotyping for the following reasons: ( i) the variations are often discovered through sequencing, which then represents a “default assay” for the SNP ( ii) in the case of highly polymorphic DNA regions, assay design may be impossible, and thus sequencing offers the opportunity to simultaneously call several SNP genotypes ( iii) sequencing equipment and technical know-how are widely available and do not require additional training or investment ( iv) in the case of difficult assays (as on genomically duplicated regions), sequencing offers additional confidence through the possibility to inspect the flanking sequence. SNPs used for simple haplotype tagging can tolerate some level of design failures, but those of particular functional relevance must be genotyped. There are scarce published data on the frequency of such design failures ( 1) in our experience, this rate is in the range of 5%-25%, depending on the method and flanking sequence. Although many specialized high-throughput methods exist, in a significant number of cases, no assay for a particular SNP can be designed due to either detection or chemistry-related problems. The genetic investigation of complex disorders requires single nucleotide polymorphism (SNP) genotyping in large patient cohorts.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
